
Plot PseudoDualSimulations
Source: R/Simulations-methods.R
plot-PseudoDualSimulations-missing-method.RdSummarize the simulations with plots.
This plot method can be applied to PseudoDualSimulations objects in
order to summarize them graphically. Possible types of plots at the moment
are:
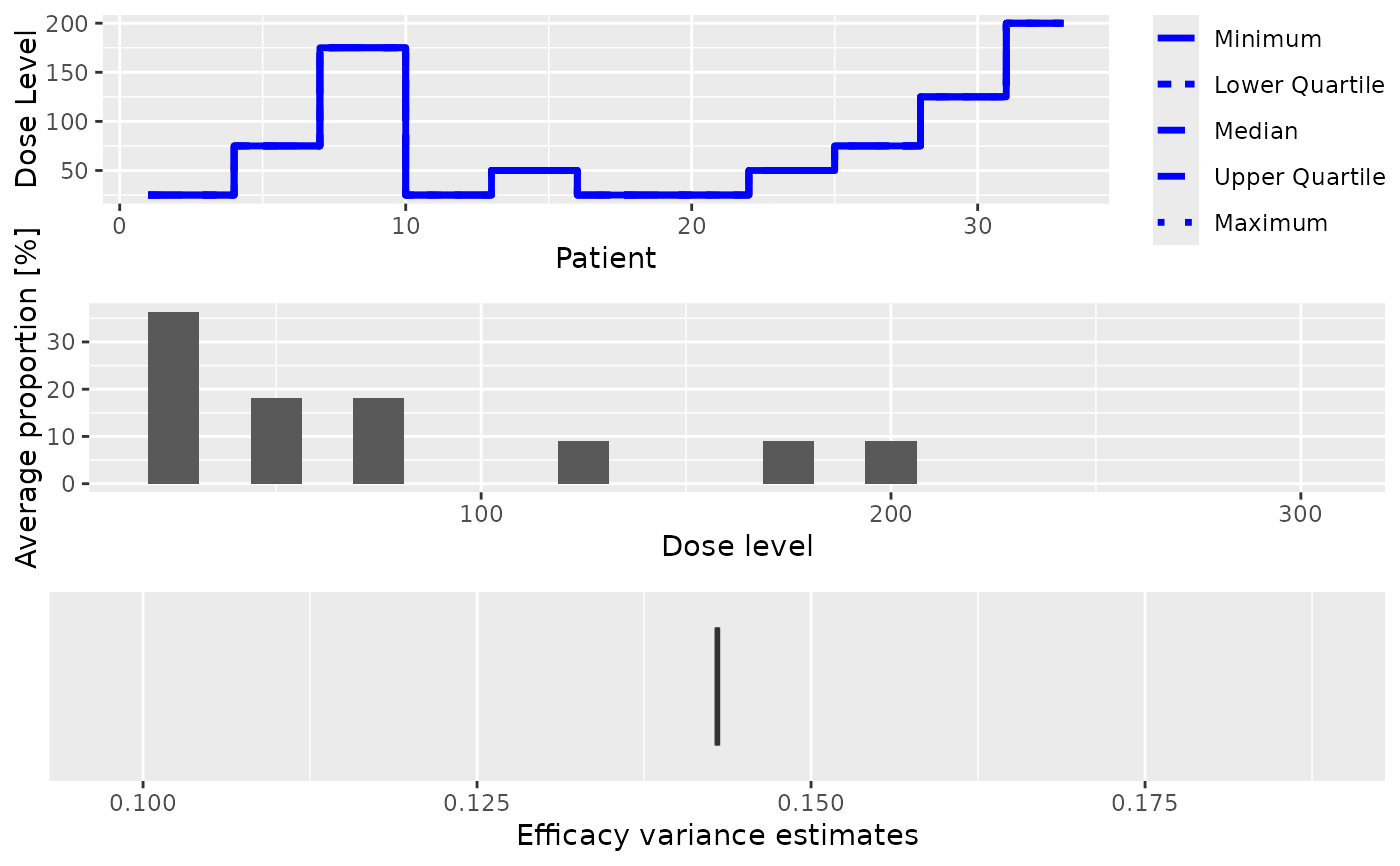
- trajectory
Summary of the trajectory of the simulated trials
- dosesTried
Average proportions of the doses tested in patients
- sigma2
The variance of the efficacy responses
You can specify one or both of these in the type argument.
Examples
# Obtain the plot for the simulation results if DLE and efficacy responses
# are considered in the simulations.
# Example to run simulations when no samples are used. The data object
# must be defined with doses >= 1:
emptydata <- DataDual(doseGrid = seq(25, 300, 25), placebo = FALSE)
# The DLE model must be of 'ModelTox' (e.g 'LogisticIndepBeta') class.
dle_model <- LogisticIndepBeta(
binDLE = c(1.05, 1.8),
DLEweights = c(3, 3),
DLEdose = c(25, 300),
data = emptydata
)
# The efficacy model must be of 'ModelEff' (e.g 'Effloglog') class.
eff_model <- Effloglog(
eff = c(1.223, 2.513),
eff_dose = c(25, 300),
nu = c(a = 1, b = 0.025),
data = emptydata
)
# The escalation rule using the 'NextBestMaxGain' class.
my_next_best <- NextBestMaxGain(
prob_target_drt = 0.35,
prob_target_eot = 0.3
)
# Allow increase of 200%.
my_increments <- IncrementsRelative(intervals = 0, increments = 2)
# Cohort size of 3.
my_size <- CohortSizeConst(size = 3)
# Stop only when 36 subjects are treated or next dose is NA.
my_stopping <- StoppingMinPatients(nPatients = 36) | StoppingMissingDose()
# Now specify the design with all the above information and starting with a
# dose of 25 (for details please refer to the 'DualResponsesDesign' example).
my_design <- DualResponsesDesign(
nextBest = my_next_best,
model = dle_model,
eff_model = eff_model,
stopping = my_stopping,
increments = my_increments,
cohort_size = my_size,
data = emptydata,
startingDose = 25
)
# Specify the true DLE and efficacy curves.
my_truth_dle <- probFunction(dle_model, phi1 = -53.66584, phi2 = 10.50499)
my_truth_eff <- efficacyFunction(
eff_model,
theta1 = -4.818429,
theta2 = 3.653058
)
# Run simulations (for illustration purpose only 1 simulation is produced).
# \donttest{
my_sim <- simulate(
object = my_design,
args = NULL,
trueDLE = my_truth_dle,
trueEff = my_truth_eff,
trueNu = 1 / 0.025,
nsim = 1,
seed = 819,
parallel = FALSE
)
# Plot the simulation results.
print(plot(my_sim))
# Example if DLE and efficacy samples are involved.
# The escalation rule using the 'NextBestMaxGainSamples' class.
my_next_best <- NextBestMaxGainSamples(
prob_target_drt = 0.35,
prob_target_eot = 0.3,
derive = function(samples) {
as.numeric(quantile(samples, prob = 0.3))
},
mg_derive = function(mg_samples) {
as.numeric(quantile(mg_samples, prob = 0.5))
}
)
# The design of 'DualResponsesSamplesDesign' class.
my_design <- DualResponsesSamplesDesign(
nextBest = my_next_best,
cohort_size = my_size,
startingDose = 25,
model = dle_model,
eff_model = eff_model,
data = emptydata,
stopping = my_stopping,
increments = my_increments
)
# Options for MCMC.
my_options <- McmcOptions(burnin = 10, step = 1, samples = 20)
# For illustration purpose only 1 simulation is produced (nsim = 1).
my_sim <- simulate(
object = my_design,
args = NULL,
trueDLE = my_truth_dle,
trueEff = my_truth_eff,
trueNu = 1 / 0.025,
nsim = 1,
mcmcOptions = my_options,
seed = 819,
parallel = FALSE
)
# Plot the simulation results.
print(plot(my_sim))
 # }
# }