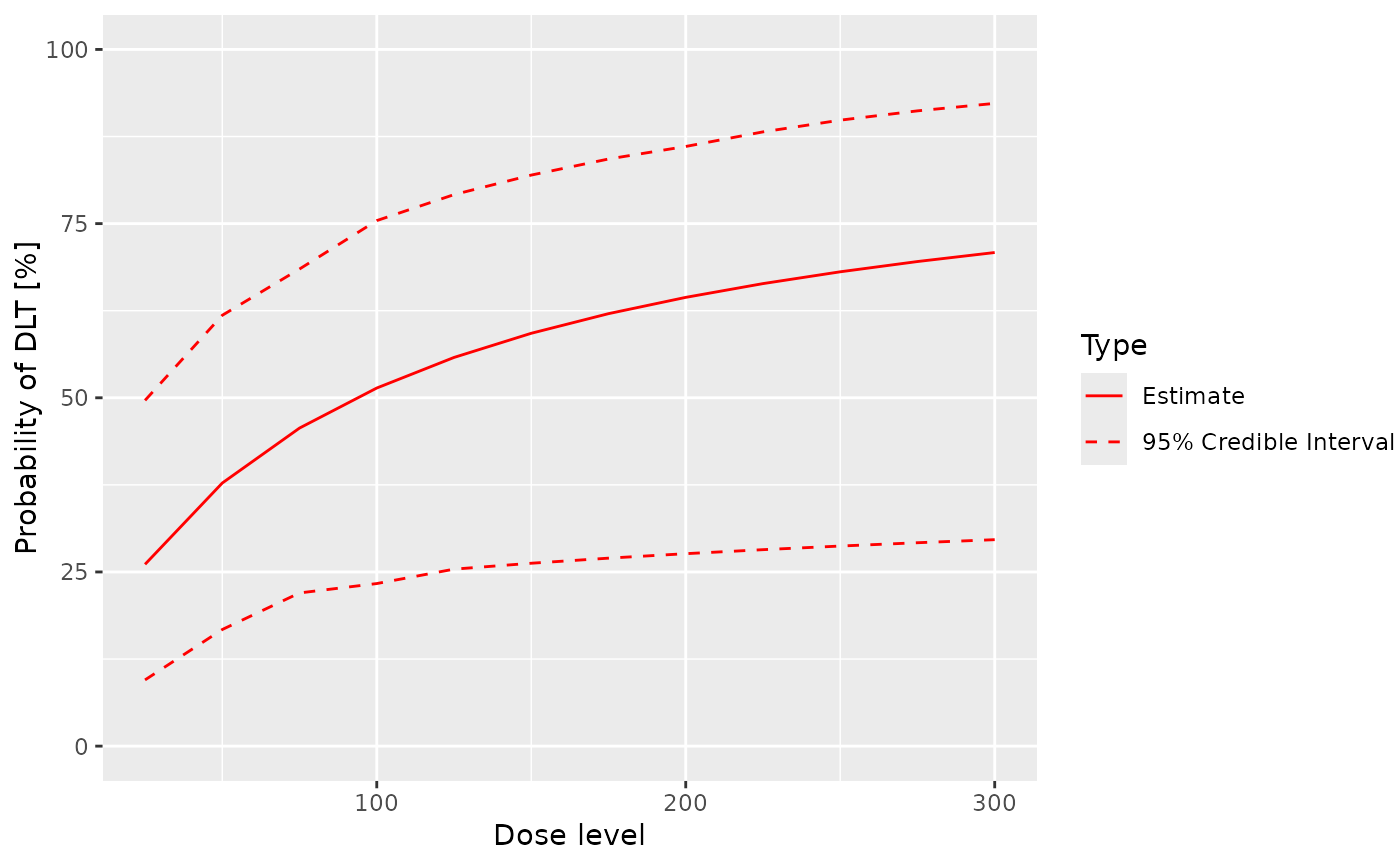
Plot the fitted dose-DLE curve using a ModelTox class model with samples
Source: R/Samples-methods.R
plot-Samples-ModelTox-method.RdPlot the fitted dose-DLE curve using a ModelTox class model with samples
Usage
# S4 method for class 'Samples,ModelTox'
plot(
x,
y,
data,
...,
xlab = "Dose level",
ylab = "Probability of DLT [%]",
showLegend = TRUE
)Value
This returns the ggplot
object for the dose-DLE model fit
Examples
## we need a data object with doses >= 1:
data <- Data(
x = c(25, 50, 50, 75, 150, 200, 225, 300),
y = c(0, 0, 0, 0, 1, 1, 1, 1),
doseGrid = seq(from = 25, to = 300, by = 25)
)
#> Used default patient IDs!
#> Used best guess cohort indices!
## plot the dose-DLE curve with samples using the model from 'ModelTox'
## class e.g. 'LogisticIndepBeta' class model
## define the model (see LogisticIndepBeta example)
model <- LogisticIndepBeta(
binDLE = c(1.05, 1.8),
DLEweights = c(3, 3),
DLEdose = c(25, 300),
data = data
)
## define the samples obtained using the 'LogisticIndepGBeta' model
## Define options for MCMC
options <- McmcOptions(burnin = 100, step = 2, samples = 200)
## (see details in 'Samples' example) samples must be of 'Samples' class
samples <- mcmc(data = data, model = model, options = options)
## plot the fitted dose-DLE curve including the 95% credibility interval of the samples
## 'x' should be of 'Samples' class and 'y' of 'ModelTox' class
plot(x = samples, y = model, data = data)